3D Printed Microscopy Chambers
Customized imaging chambers can be 3D printed using FDM or SLA printing technology, and attached to a #1.5 thickness coverglass to allow for high-resolution imaging. When prepared properly, these chambers show no cytotoxicity or other signs of altered cell behaviour, even in cells cultured for multiple days. These are the chamber designs used in the Heit Lab, and are based on our publication in Biochemistry and Cell Biology[1], and on the Mod3D preprint[2] by the Truant Lab.
In addition to directly printing chambers, 3D printing can also be used to make moulds for casting PDMS chambers, as described in our publication in Biochemistry and Cell Biology[1].
Chamber Designs
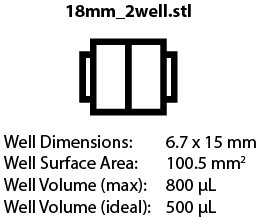
Chambers are available to fit 18 mm x 18 mm and 24 mm x 50 mm coverslips, with different well arrangements. We are still testing some of these designs, but the images below will link to the corresponding .STL file once designs are finalized.
 |
 |
 |
|---|---|---|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
FDM (Filament) Printing
FDM printing is the easiest in terms of printing and post-printing assembly, but it is more difficult to get a good seal with the coverslip than with DLP (resin) printing. It is important to use a good quality filament free of potentially cytotoxic or fluorescent compounds - we recommend a black or unpigmented, FDA-food grade certified PLA or PETG filament such as those sold by Filaments.ca.

Recommended Print Settings:
- 0.4 mm diameter nozzle
- 100% infill
- 0.1 mm layer height
- Print speed/acceleration set for high quality prints (varies with printer)
- Ironing:
- Top-most surface for chambers (this helps with the bond to the coverslip)
- None on Lids
- All top surfaces for alignment guides
- No supports
FDM Printing
- Layout your desired prints using PrusaSlicer (or your preferred slicing software). Be certain to position the prints such that the surface that will contact the coverslip faces upwards. The numbers and letters on the chamber will be upside-down if the chamber is positioned correctly.
- Configure the print settings as above and slice.
- Carefully remove the print from the print bed and wash for 15 min in distilled water.
- Dry chamber before assembly (see below).
DLP (Resin) Printing

DLP (resin) printing provides superior resolution and a flatter surface for bonding the coverslip to the chamber. However, the prints must be treated carefully to remove any residual polymer or photocatalyst, as these are highly cytotoxic. At this time the only verified resin for this purpose is eSun's eResin-PLA Bio-Photopolymer Resin (Black).
Recommended Print Settings:
- Maximum x/y resolution
- No anti-aliasing
- 0.05 mm layer height
- Exposure times need to be optimized for individual instruments, but for our Elegoo Mars 3:
- 4 burn-in layers @ 60 s
- 0 transition layers
- 4 s light-off delay
- 6 mm lift
- 0.4 mm/m lift speed
- 150 mm/m retract speed
- Normal layers:
- 0.05 mm layer height
- 8.5 s exposure
- 0 mm lift distance
- 65 mm/m lift speed
- 150 mm/m retract speed
- 0 light-off delay
- 4 burn-in layers @ 60 s

DLP Printing:
- Layout prints as desired in Lychee Slicer or Chitubox (or your preferred slicing software). Be certain to position the prints such that the surface that will contact the coverslip face upwards. The numbers and letters on the chamber will be upside-down if the chamber is positioned correctly.
- Rotate the prints 40° on the short axis (e.g. objects are rotated along the long axis, image to right).
- Use autosupports to lift and generate supports for the chambers. A minimum of 3 mm lift is recommended.
- The prints are soft at this point, so treat them gently for the next few steps.
- Carefully remove the print from the print bed and place in a wash container of isopropyl alcohol. It is fine to use isopropyl that has been used previously for this step.
- Using flat cutters carefully remove all supports.
- Place the chambers in a dedicated cleaning container, coverslip-contact side up. Do not stack the prints. Cover compeltely with clean (previously unused) isopropyl alcohol. Wash with gently shaking for 20 minutes.
- Carefully remove the chamber and place in the curing station, coverslip-contact side up. Position the chambers such that they do not contact each other. Cure for 20 minutes, then flip the parts and cure an additional 20 minutes. Do not crowd parts during this step - it is better to cure in multiple batches than it is to have shadowed regions that do no cure.
Assembly
Assembly is the same whether the chambers are printed via DLP or FDM.
- Acid wash coverslips in advance of assembly day, ensuring they are completely dry and dust free before assembly.
- Place a coverslip into the bottom of the alignment guide.
- Place a small drop of SS-433T silicone glue (Silicone Solutions, OH, USA) on a phenolic sheet, and using a roller, spread it into a thin (~0.5 mm) thick layer that is larger than your chamber. Note that PDMS can be used as well (10:1 polymer:catalyst), but the bond will not be as strong.
- Press the coverslip-contact part of the chamber into the glue, and check for even glue across the entire sealing surface. If the whole surface is not coated, press the chamber into the glue again. Repeat until all coverslip-contact surfaces are whetted with glue.
- Put the chamber (glue-side down) into the alignment guide and press onto the coverslip for 10 seconds.
- Carefully invert the alignment guide and remove the chamber.
- Check the seal - a good seal will appear wet; any dry-appearing areas are not glued. If you see a dry area, gently press that part of the chamber with your finger. If a good seal cannot be made, remove the coverslip and repeat steps 2-6 with a new coverslip. This step is critical, as poorly glued chambers will leak.
- Transfer the completed chambers to a humidified, 37°C incubator for 24 hours to cure.
References
- ↑ 1.0 1.1 Tepperman A, Zheng DJ, Taka MA, Vrieze A, Le Lam A, Heit B. Customizable live-cell imaging chambers for multimodal and multiplex fluorescence microscopy. Biochem Cell Biol. 2020 Oct;98(5):612-623. doi: 10.1139/bcb-2020-0064. Epub 2020 Apr 27. PMID: 32339465. Link to Paper. Link to Preprint.
- ↑ C. Barba Bazan, S. Goss, C. Peng, N. Begeja, CE. Suart, K. Neuman, Ray Truant. Mod3D: A Low-Cost, Flexible Modular System of Live-Cell Microscopy Chambers and Holders. bioRxiv 2021.10.18.462400; doi: https://doi.org/10.1101/2021.10.18.462400